Spatial Datasets
Spatial transcriptomic datasets can be visualized and analyzed in rakaia. The preprocessing steps for spatial datasets are slightly different from antibody-based imaging datasets, and will also vary slightly across different technologies.
Required format
Users will likely need to use a spatial analysis library to prepare their data for import, such as spatialdata or scanpy. Import of raw instrument outputs, such as 10x output bundles, are not directly supported.
IMPORTANT: If importing with a zarr directory path, users should not add a forward or backward trailing slash (/ or \) to the path, or they may receive an import error like the following:

All spatial datasets that contain marker expression need to be imported into rakaia as one of two formats:
- as a
spatialdatazarrdirectory, created using the appropriate technology-specific reader fromspatialdata. This currently supports 10x Visium, Visium HD, and Xenium. More information onspatialdatafile reading can be found here - as an Anndata object with the file extension
.h5adSpecifically, the Anndata object must have aspatialarray in theobsmslot that contains both the x and y coordinates for each spatial measurement, whether it corresponds to a cell, spot, etc. Libraries such asscanpyandsquidpyare Python libraries that have provide readers for raw spatial data into this format. These libraries will be referenced in the article below.
rakaia currently expects every spatial dataset sample or slide (e.g. Visium slide/Xenium sample run) to be loaded as its own .h5ad file. If users have multiple samples or slides, each with a unique set of spots or in-situ transcripts, the dataset should be divided into multiple Anndata objects and exported as individual files.
rakaia also enables multi-slide datasets with 10x Visium through spatialdata + zarr (more info below).
Importing multiple zarr stores per session
From rakaia 0.26.0 and later, users can import multiple zarr stores into one session provided that the biomarker panel is the same across all samples. For example, with the following data directory structure:
The user can import all 4 spatial ROIs into the session by simply importing from the parent filepath:
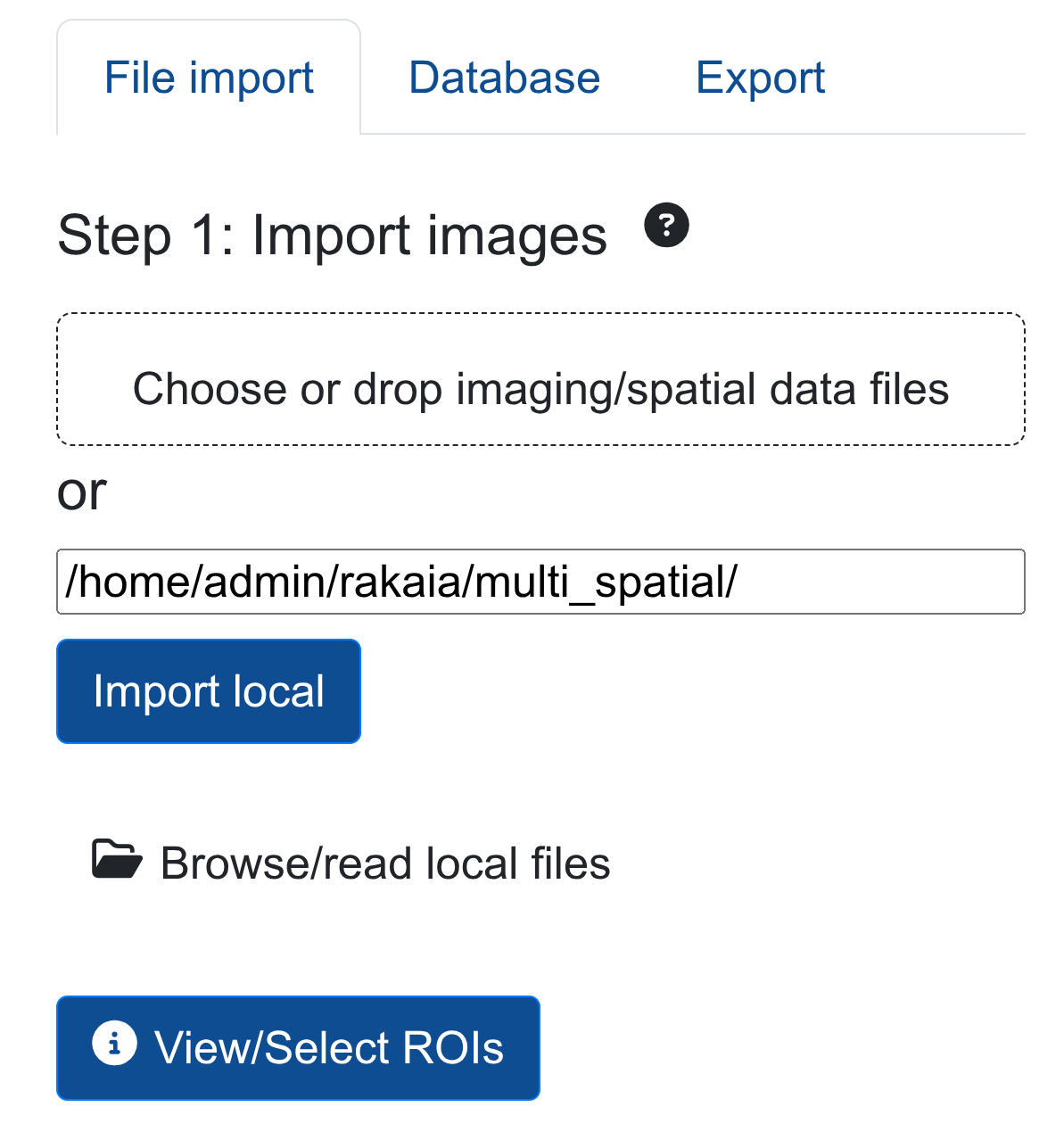
This will parse all 4 zarr directories into the session, allowing for efficient multi-sample or multi-slide spatial analysis.
Spot-based assays: 10X Visium V1, V2
Spot-based spatial technologies such as the 10X Visium Spatial Gene Expression profile transcript counts summarized at the spot level. The Visium technology is capable of profiling tens of thousands of markers per spot, providing comprehensive spatial context of the transcriptome.
Raw Visium data (from the space ranger standard directory output as described here) should be read using either read_visium function from either scanpy or squidpy. Below is a minimal example showing how the data can be preprocessed and exported into a compatible file format:
import squidpy as sq
import os
# specify the input directory with the outs subdir
input_dir = "/path_to_visium_raw/outs/"
adata = sq.read_visium(input_dir)
adata.var_names_make_unique()
# specify the output file as an anndata object
out_anndata = "/output_dir/visium.h5ad"
adata.write_h5ad(out_anndata)
The output h5ad Visium file should typically not be larger than 200-300mb. If the size significantly exceeds this range (1-2 GB or larger), then it is likely that the user has cached a full-sized WSI (i.e. H & E) in the file. Caching these large images will result in significant performance slowdowns in rakaia. To avoid this, users should ensure that the uns slot in the object is cleared of any fields except the required scalefactors slot:
adata.uns = {'spatial': {str(list(adata.uns['spatial'].keys())[0]): {
'scalefactors': adata.uns[
'spatial'][list(adata.uns['spatial'].keys())[0]]['scalefactors']}}}
This retains the scale factors required for Visium to render in rakaia, while removing any additional slot caches that rakaia does not use.
When the Anndata object is read into rakaia, the spot size and region dimension will automatically be computed from the spatial coordinates and scale factors:
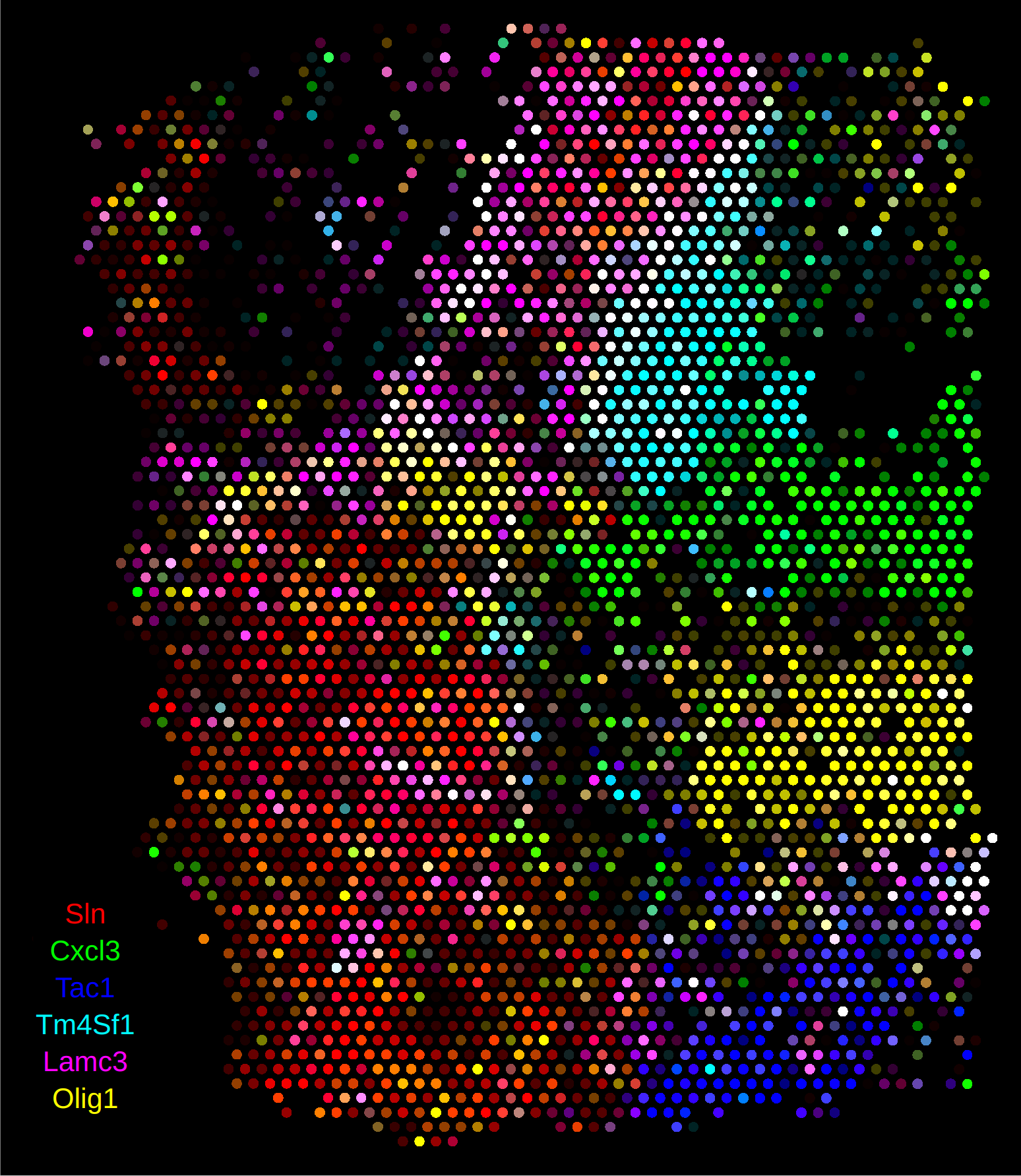
10x Visium with spatialdata + zarr
Visium assays read through a spatialdata zarr directory can have multiple samples/slides. Every unique sample or slide should have its own shape frame in the shapes slot. The spatialdata_io.visium reader should be used to generate the zarr store (More information can be found here)
Non-spot based assays
Non spot-based assays behave differently in rakaia as the user has more flexibility over the visualization parameters. Specifically, this means that the user may specify a unique visualization size for the data in the image, which isn't supported with spot-based assays because the spot scaling factors are computed automatically from the input data.
Binned expression assays: 10x Visium HD
The HD version of 10X Visium differs slightly from the spot-based technology used in either V1 or V2. Instead of using circular spots that have gaps among them, HD offers tiled, barcoded squares without gaps, as described here. This results in data that can be binned at summarized at three different micron resolutions: 2, 8, and 16.
Currently, the only supported reader in Python for HD datasets is the spatialdata HD reader. The reader generates aggregate expression profiles for each micron resolution, and each of these profiles can be exported as an Anndata file for visualization in rakaia. This example notebook here shows an example of how to export these binned profiles. each bin can then be imported into rakaia as a separate ROI with the same set of marker genes, but with different dimensions and resolution.
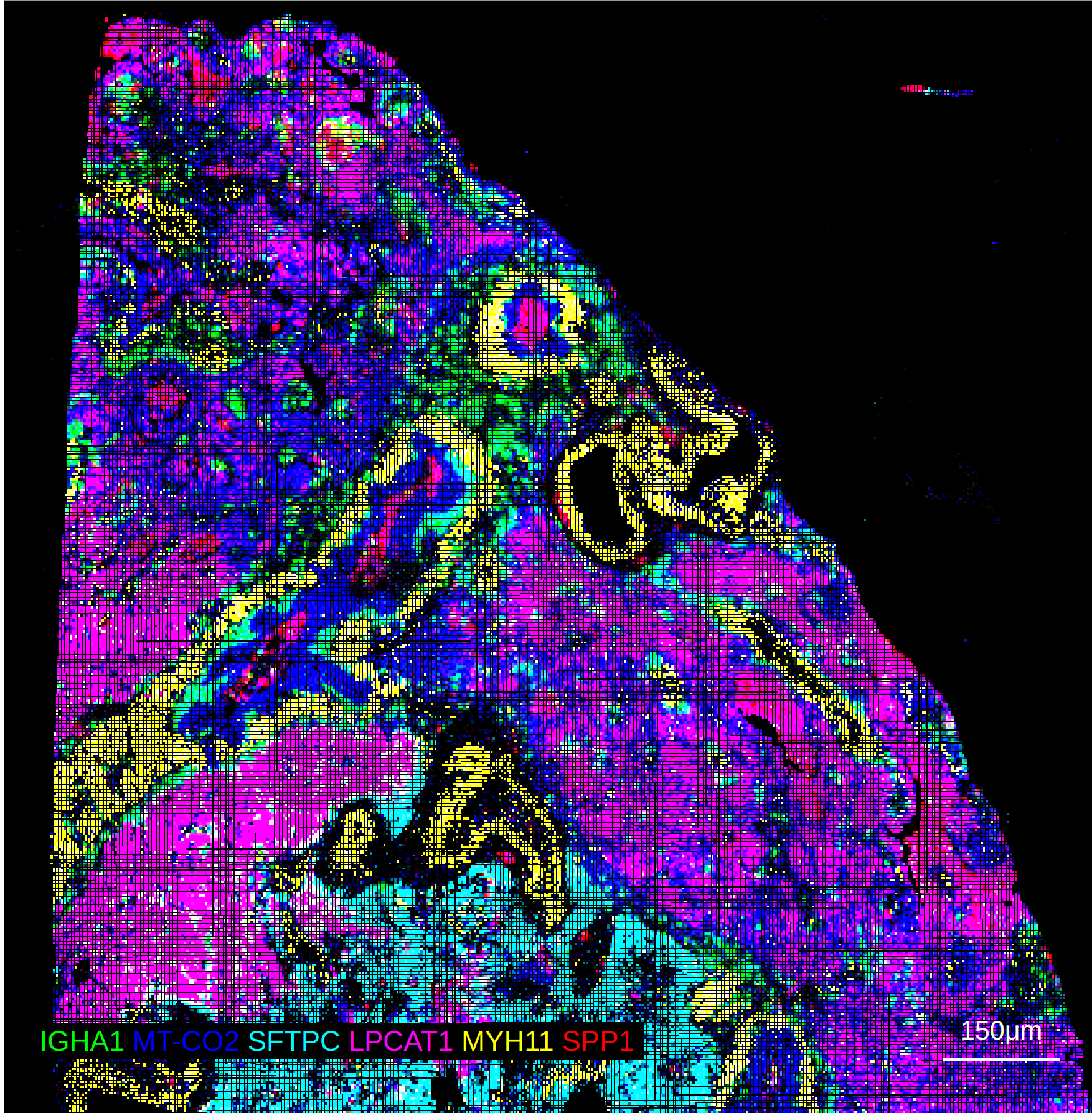

Visium HD with spatialdata + zarr
Visium HD assays read through spatialdata + zarr will be split into multiple ROIs based on the bin size. The zarr store should have a matched shape + table slot for every bin size with the following table keys: 'square_002um', 'square_008um', 'square_016um'.The spatialdata_io.visium_hd reader should be used to generate the zarr store (More information can be found here)
rakaia will output only the bins for 8 and 16um, as the 2um bin size is both too sparse and large to effectively render.
From rakaia v0.25.0, the segmentation masks for Visium HD will be available for the bin sizes above. However, due to the binning resolution, the masks will not retain the same shape and resolution as they would appear in the Visium HD cell segmentation summary, so they should be used more for positional inference or object ID annotation rather than accurate spatial projection.
10X Xenium In-situ Expression�
The 10X Xenium platform differs slightly from the assays above as it profiles in-situ transcript counts, and also supports segmentation and the overlay of object masks. The notebook example here provides a complete example of reading the data into a spatialdata object and exporting both the expression Anndata object as well as the separate cell segmentation mask as a tiff.
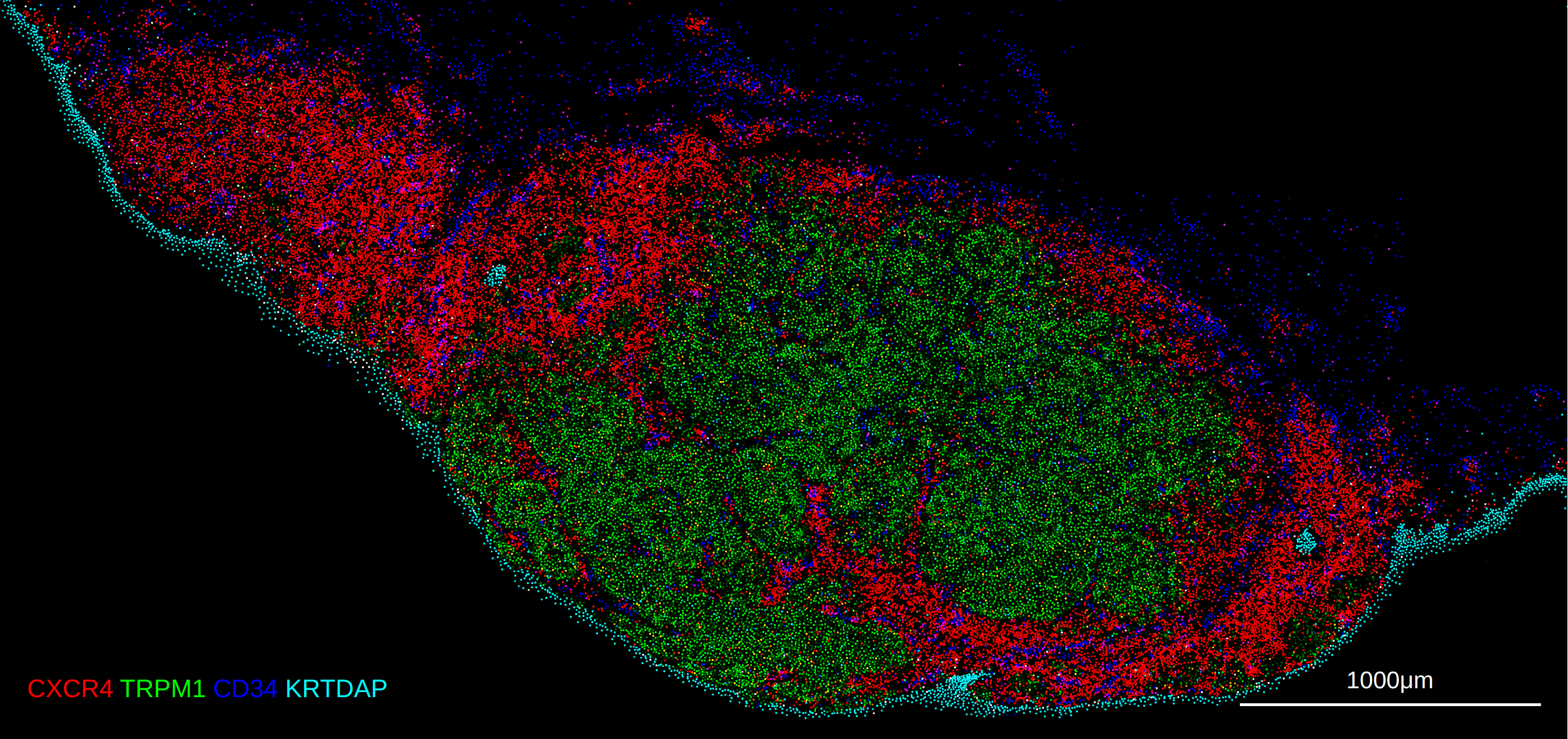
Xenium with spatialdata + zarr
Xenium read through spatialdata + zarr should be a single slide/ROI per zarr directory. The zarr store will parse for cell or nucleus segmentation objects in the following shapes slot keys: 'cell_boundaries', 'cell_circles', 'nucleus_boundaries', and will import the cell boundary mask. These slots are strictly optional; if no segmentation results are found, then rakaia will parse the spatialdata store as a default expression Anndata (see below)
Users are encouraged to create a "lean" spatialdata object store with the code template below, where large-memory data structures that rakaia currently does not import are ignored; this will benefit the user by minimizing memory usage when creating spatialdata stores, and make them faster to read in rakaia sessions:
sdata = xenium(xenium_raw_output_dir, morphology_focus=False, cells_boundaries=True,
morphology_mip=False, aligned_images=False,
nucleus_boundaries=False, nucleus_labels=False,
cells_table=False, cells_labels=False, transcripts=False,
n_jobs=8)
In the template above, the resulting sdata object will contain only the cell aggregated probe/transcript counts as well as the cell boundary mask, which are both rendered by rakaia. Other large data stores such as high-res morphology images, nuclei-related outputs, and individual transcript counts are currently not used by rakaia, so withholding them from the spatialdata object will make reading large Xenium samples much more memory-friendly and faster.
Rakaia supports both the spatialdata_io.xenium and sopa.io.xenium API for spatialdata zarr stores.
Setting marker sizes for visualization (non spot-based)
The non spot-based technologies above support custom user marker sizes in rakaia. This means that the marker can be visually enlarged or minimized down to a minimum size of 1 pixel in the viewer. This allows the expression to appear more granular/minimal or more extensive/uniform throughout the ROI as the user desires.
Below are some recommended marker sizes for the different technologies listed above. The marker size can be changed under Additional application settings -> Appearance -> Custom spatial marker radius.
-
10X Visium HD: 2-3. A marker size of 2 will generally reveal pixel gaps between areas of expression, with 3 removing those gaps and providing a more uniform, albeit maybe slightly blurred, expression visualization. Generally, marker sizes of 4 of greater will cause expression to overlap erroneously.
-
10X Visium: 2-4. The marker size should be set based on the presence of an overlaid segmentation mask. Without using a segmentation mask, larger marker sizes up to 4 will make the expression more visually appealing at a global resolution, but could result in markers that "spill" out of the segmentation mask. When using a mask, values of 2-3 allow the marker to appear in the centroid of the cell mask while also allowing the expression to appear at the global resolution.
Setting a larger marker size increases the possibility of seeing overlapping or fragmented expression points in 10X Xenium assays. This is particularly noticeable in dense tissue where cells may be close together, or areas where it is difficult to clearly segment cell boundaries. Additionally, filtering transcripts using a lower bound may cause visual fragmentation of points, as a portion of the overlapping expression point may be filtered out. In these instances, users may want to reduce the marker size incrementally until the markers no longer touch/overlap.
The default marker size for spatial assays is set at 4.
Processing and filtering considerations
When users are filtering or performing some preprocessing of the spatial data assays above where cells and/or spatial locations are removed, special care should be taken that the other slots in the spatialdata object such as mask polygons, etc. are also filtered to match the expression data. While Rakaia does explicitly check for this when parsing from zarr directories, users should perform these additional checks to ensure that all data slots in a spatialdata object remain aligned.
As an example, with 10x Xenium, filtering out cells with low cell counts causes a mismatch between the cell-level probe counts in sdata.tables['table'] and sdata.shapes['cell_boundaries']. The following code snippet helps to ensure that these remain matched:
expr = sdata.tables['table']
cells = sdata.shapes['cell_boundaries']
# filter out cells with fewer than 25 probe counts
# Compute total counts per cell
total_counts = np.array(expr.X.sum(axis=1)).flatten()
# Subset AnnData (keep cells with >= 25 counts)
expr = expr[total_counts >= 25].copy()
# retain cells only in the new expression data
cells = cells.loc[cells.index.isin(expr.obs['cell_id'])]
sdata.tables['table'] = expr
sdata.shapes['cell_boundaries'] = cells
Other spatial datasets
Additional spatial technologies may be inherently supported in rakaia provided that they follow the input data format as described above (minimally, that spatial coordinates for pixel locations are provided in spatial array in the obs slot).
Additional examples of spatial datasets that can be rendered in rakaia include the following from the squidpy spatial technology tutorials:
-
Vizgen: https://squidpy.readthedocs.io/en/stable/notebooks/tutorials/tutorial_vizgen.html
-
4i: https://squidpy.readthedocs.io/en/stable/notebooks/tutorials/tutorial_fouri.html
-
Slide-seq: https://squidpy.readthedocs.io/en/stable/notebooks/tutorials/tutorial_slideseqv2.html
For all tutorial datasets above, the data are imported and analyzed in Anndata format, consistent with the required format for rakaia. The adata object referenced in the tutorials can be exported as files with the .h5ad format.
Non-10x spatial assays in spatialdata zarr stores
If rakaia is not able to parse out a 10x compatible assay from the zarr store, by default, it will look for a table in the tables slot with the table key, and import that as an Anndata-formatted object with spatially resolved expression values at x and y coordinates. This is equivalent to the Anndata specified format under Required format. If this named key slot does not exist, then it will inform the user of key error and no data will be imported.
rakaia features not available to spatial datasets
Spatial datasets have orders of magnitude more variables and markers than IMC datasets (tens of thousands of markers as opposed to 40-50 antibodies), so the lazy loading features behave differently for these technologies. This means that certain features that can be applied to an entire ROI for IMC cannot be used for spatial analysis due to memory and time constraints in processing all dataset markers:
- The channel/marker tile gallery is not supported for as generating a thumbnail preview for thousands of markers would be prohibitively time-consuming
- Both marker correlation and in-app marker quantification can be performed only on markers that are in the current canvas, as these are the only variables that have been loaded into memory at a given point in analysis
Troubleshooting
Minimum package versioning
To ensure that zarr stores can be read reliably by rakaia, and to avoid any backwards incompatibility issues with zarr data encodings, users should use the following packages or newer when generating zarr directories:
anndata==0.11.0
spatial_image==1.2.1
spatialdata==0.4.0
spatialdata-io==0.3.0
spatialdata-plot==0.2.11
xarray==2024.10.0
xarray-dataclasses==1.9.1
xarray-schema==0.0.3
xarray-spatial==0.4.0
This will help to avoid errors such as
anndata._io.specs.registry.IORegistryError: No read method registered for IOSpec(encoding_type='null', encoding_version='0.1.0') from <class 'zarr.core.Array'>. You may need to update your installation of anndata.
which can occur when attempting to read zarr stores that were generated with older package versions.